Introduction
The world is moving rapidly into food crisis due to extreme weather, shortage of farmland, and population explosion. And glucose is the basic material of food produced by green plants using the natural photosynthesis (NPS)1. However, the NPS suffers from low-energy efficiency (~1%) and is greatly limited by soil, climate, water, and labor2,3. It is reported that food demand is expected to increase by 70% by 20504, which makes an extremely urgent demand for a scientific solution beyond NPS for food production. Recently, scientists have figured out an excellent alternative to NPS, which is called artificial photosynthesis (APS), for the energy-rich chemicals production. It mimics the green plants to produce energy-rich materials and fuels but using man-made nanomaterials and engineered photosynthetic reactors. Compared with the NPS, the APS usually presents much higher efficiency (~12%) and simplicity5. Although the APS has attracted substantial research interest, most of efforts have been devoted to the light-dependent reactions for hydrogen generation and water splitting6,7, microbial growth and biofuel production8,9,10,11,12, and photocatalytic cofactor regeneration13,14,15,16. The exploration of light-independent reactions for food production is worth more attentions. Nevertheless, it is found that the light-independent reactions, though crucial for glucose production from CO2, are more challenging for the artificial replication, since they involve multi-step enzymatic reactions (Fig. 1a).
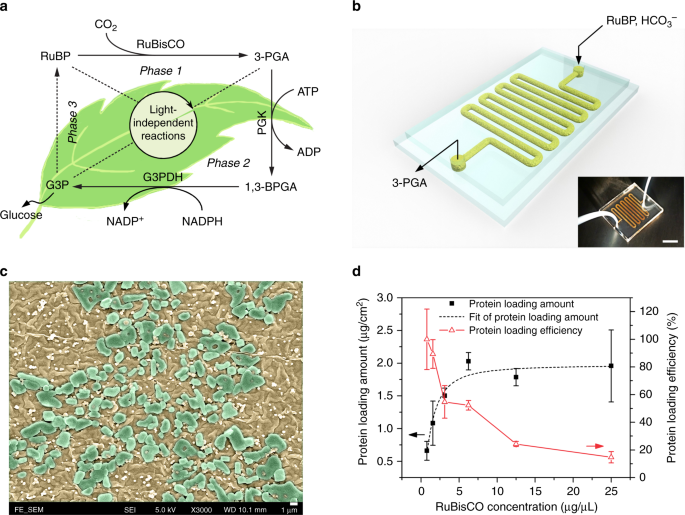
Design and characterization of RuBisCO immobilized microfluidic reactors (RIMRs). a Scheme of light-independent reactions of NPS: Phase 1: Carbon fixation starts with ribulose 1,5-bisphosphate (RuBP) and uses the enzyme RuBisCO to fix CO2 into 3-phosphoglycerate (3-PGA); Phase 2: Reduction reaction uses adenosine triphosphate (ATP), nicotinamide adenine dinucleotide phosphate (NADPH) and the enzyme phosphoglycerate kinase (PGK) and glyceraldehyde 3-phosphate dehydrogenase (G3PDH) to reduce 3-PGA into glyceraldehyde 3-phosphate (G3P), two of which can form the end product glucose; Phase 3: RuBP regeneration from G3P using up to 9 steps of enzymatic reactions. b Three-dimensional diagram and the photograph (inset) of the RIMRs, the scale bar of the inset is 1 cm. c SEM image of the inner surfaces of RIMRs. Flat and smooth PDMS becomes rough and is covered by PDA nanoparticles after PDA modification (brownish color). Large blocks are the immobilized RuBisCO (green blocks). The scale bar is 1 μm. d Protein-loading amount and protein-loading efficiency as a function of the concentrations of injected RuBisCO to find the optimal RuBisCO concentration for further experiments. Error bars represent the standard deviations from three independent experiments. Source data are provided as a Source Data file.